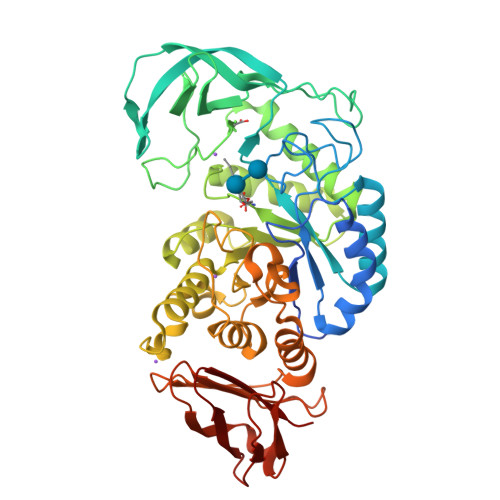
Precision Activity-Based alpha-Amylase Probes for Dissection and Annotation of Linear and Branched-Chain Starch-Degrading Enzymes.
Pickles, I.B., Chen, Y., Moroz, O., Brown, H.A., de Boer, C., Armstrong, Z., McGregor, N.G.S., Artola, M., Codee, J.D.C., Koropatkin, N.M., Overkleeft, H.S., Davies, G.J.(2025) Angew Chem Int Ed Engl 64: e202415219-e202415219
- PubMed: 39601378
- DOI: https://doi.org/10.1002/anie.202415219
- Primary Citation of Related Structures:
9FYZ, 9FZ0, 9FZ2, 9FZ3 - PubMed Abstract:
α-Amylases are the workhorse enzymes of starch degradation. They are central to human health, including as targets for anti-diabetic compounds, but are also the key enzymes in the industrial processing of starch for biofuels, corn syrups, brewing and detergents. Dissection of the activity, specificity and stability of α-amylases is crucial to understanding their biology and allowing their exploitation. Yet, functional characterization lags behind DNA sequencing and genomics; and new tools are required for rapid analysis of α-amylase function. Here, we design, synthesize and apply new branched α-amylase activity-based probes. Using both α-1,6 branched and unbranched α-1,4 maltobiose activity-based probes we were able to explore the stability and substrate specificity of both a panel of human gut microbial α-amylases and a panel of industrially relevant α-amylases. We also demonstrate how we can detect and annotate the substrate specificity of α-amylases in the complex cell lysate of both a prominent gut microbe and a diverse compost sample by in-gel fluorescence and mass spectrometry. A toolbox of starch-active activity-based probes will enable rapid functional dissection of α-amylases. We envisage activity-based probes contributing to better selection and engineering of enzymes for industrial application as well as fundamental analysis of enzymes in human health.
Organizational Affiliation:
University of York, York Structural Biology Laboratory, Department of Chemistry, UNITED KINGDOM OF GREAT BRITAIN AND NORTHERN IRELAND.