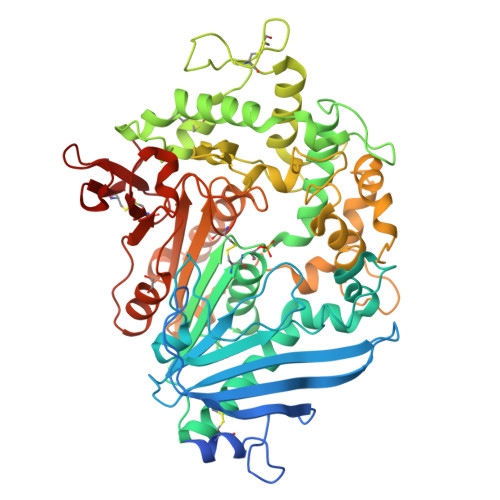
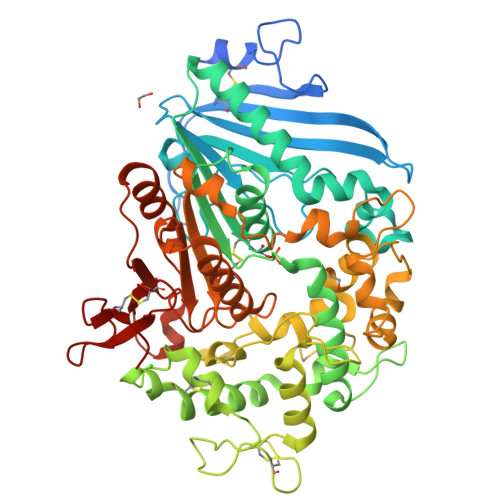
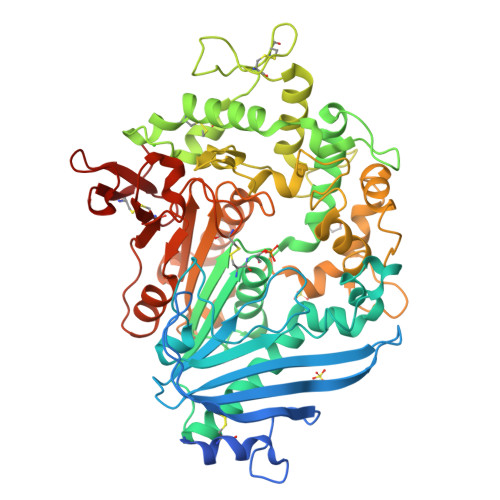
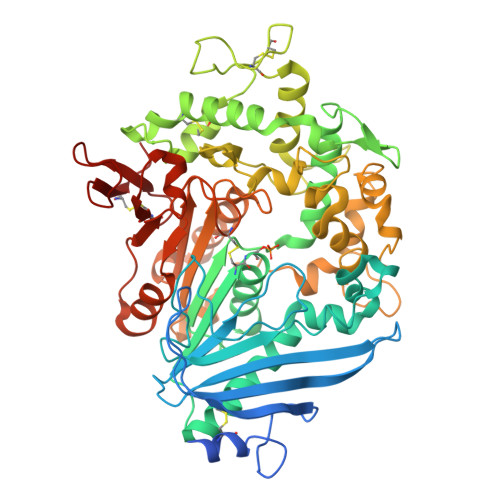
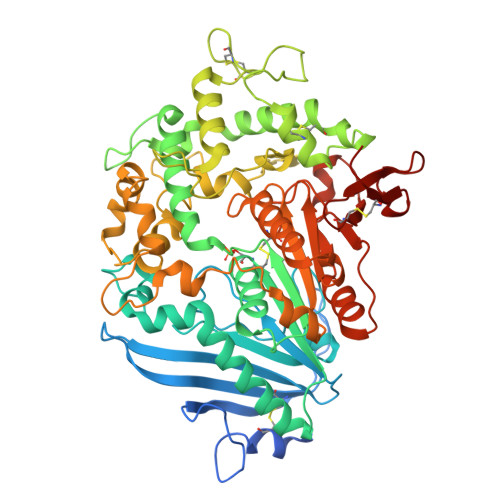
Increasing the Soluble Expression and Whole-Cell Activity of the Plastic-Degrading Enzyme MHETase through Consensus Design.
Saunders, J.W., Damry, A.M., Vongsouthi, V., Spence, M.A., Frkic, R.L., Gomez, C., Yates, P.A., Matthews, D.S., Tokuriki, N., McLeod, M.D., Jackson, C.J.(2024) Biochemistry 63: 1663-1673
- PubMed: 38885634
- DOI: https://doi.org/10.1021/acs.biochem.4c00165
- Primary Citation of Related Structures:
8EKG - PubMed Abstract:
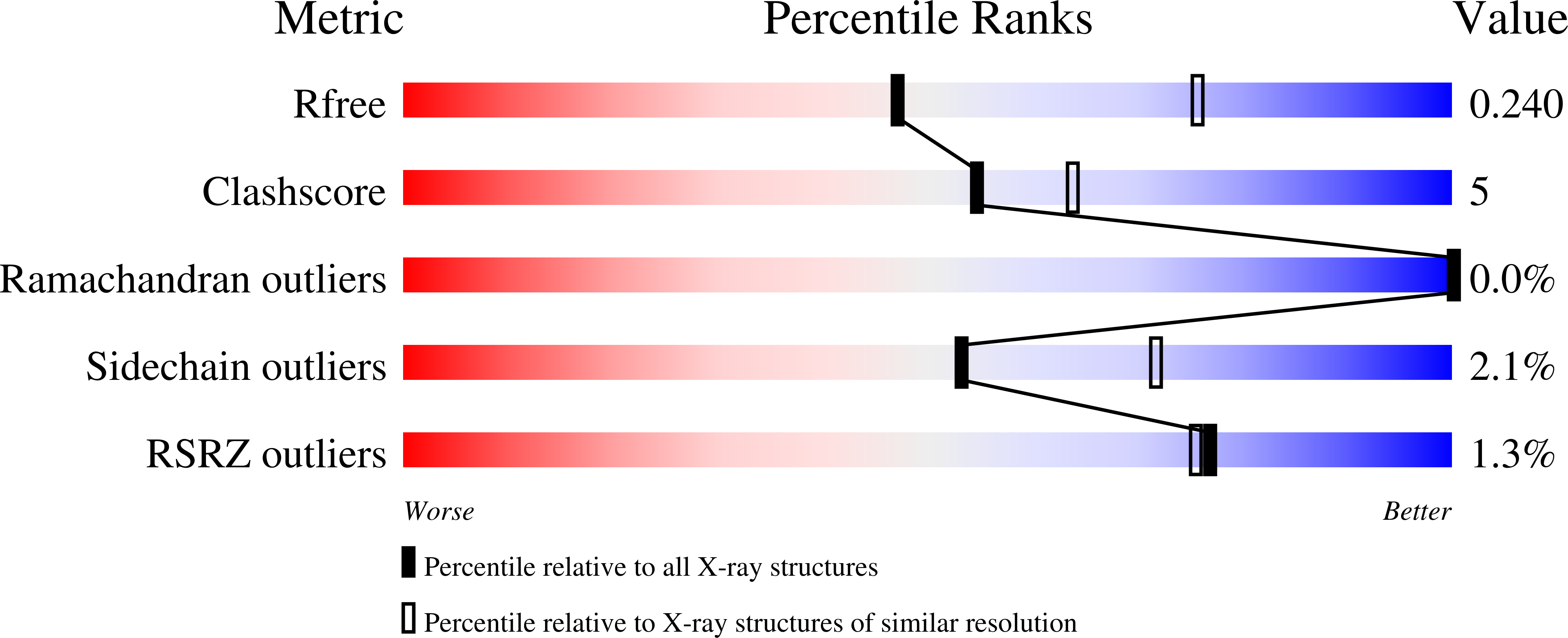
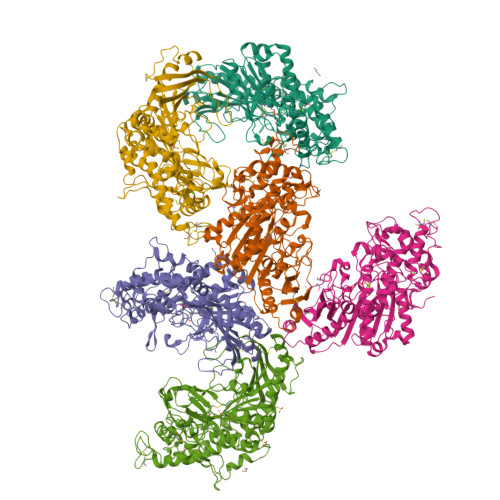
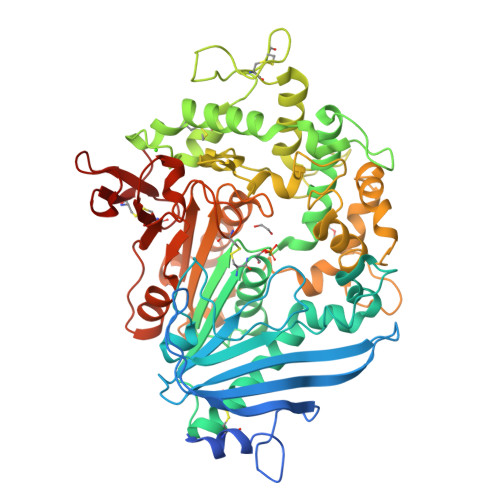
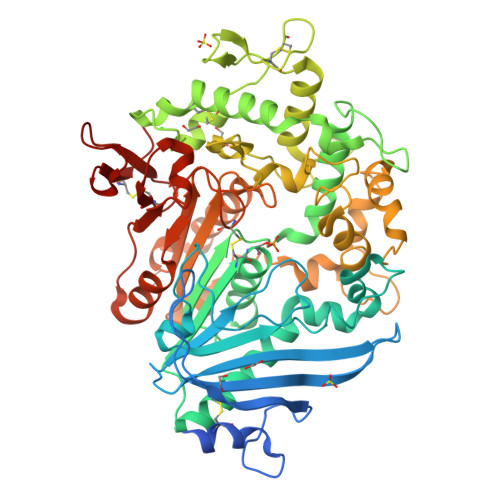
The mono(2-hydroxyethyl) terephthalate hydrolase (MHETase) from Ideonella sakaiensis carries out the second step in the enzymatic depolymerization of poly(ethylene terephthalate) (PET) plastic into the monomers terephthalic acid (TPA) and ethylene glycol (EG). Despite its potential industrial and environmental applications, poor recombinant expression of MHETase has been an obstacle to its industrial application. To overcome this barrier, we developed an assay allowing for the medium-throughput quantification of MHETase activity in cell lysates and whole-cell suspensions, which allowed us to screen a library of engineered variants. Using consensus design, we generated several improved variants that exhibit over 10-fold greater whole-cell activity than wild-type (WT) MHETase. This is revealed to be largely due to increased soluble expression, which biochemical and structural analysis indicates is due to improved protein folding.
Organizational Affiliation:
Research School of Chemistry, Australian National University, Canberra, ACT 2601, Australia.