Molecular insights and rational engineering of a compact CRISPR-Cas effector Cas12h1 with a broad-spectrum PAM.
Zheng, W., Li, H., Liu, M., Wei, Y., Liu, B., Li, Z., Xiong, C., Huang, S., Hu, C., Ouyang, S.(2025) Signal Transduct Target Ther 10: 66-66
- PubMed: 39955288
- DOI: https://doi.org/10.1038/s41392-025-02147-5
- Primary Citation of Related Structures:
8Y9L, 8Y9M, 8Y9N - PubMed Abstract:
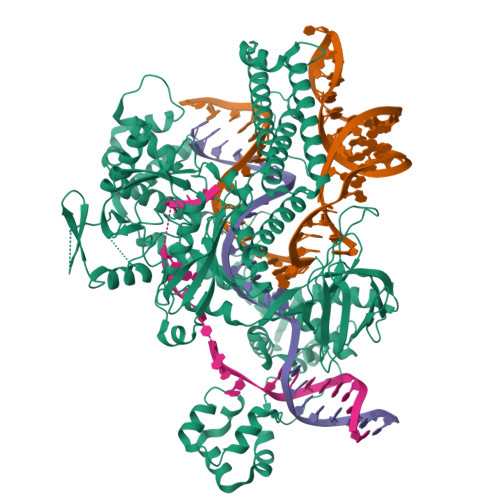
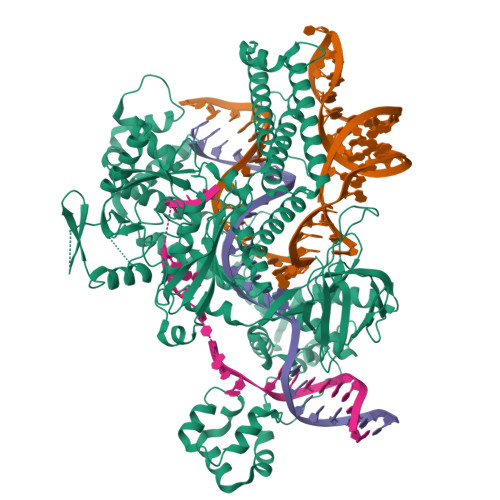
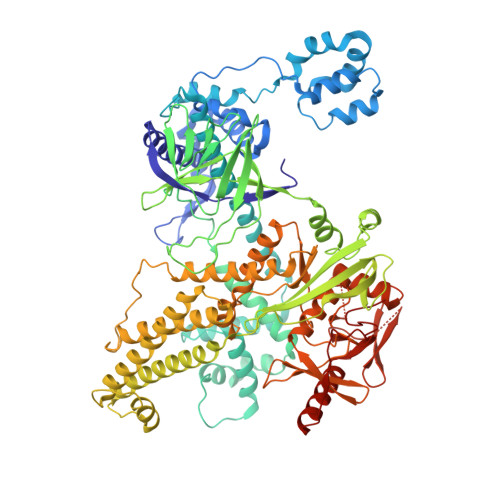
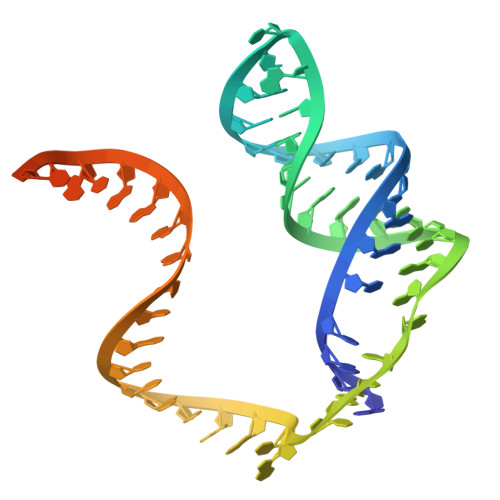
Cas12h1 is a compact CRISPR-associated nuclease from functionally diverse type V CRISPR-Cas effectors and recognizes a purine-rich protospacer adjacent motif (PAM) distinct from that of other type V Cas effectors. Here, we report the nickase preference of Cas12h1, which predominantly cleaves the nontarget strand (NTS) of a double-stranded DNA (dsDNA) substrate. In addition, Cas12h1 acts as a nickase in human cells. We further determined the cryo-EM structures of Cas12h1 in the surveillance, R-loop formation, and interference states, revealing the molecular mechanisms involved in the crRNA maturation, target recognition, R-loop formation, nuclease activation and target degradation. Cas12h1 notably recognizes a broad 5'-DHR-3' PAM (D is A, G, or T; H is A, C, or T; R is A or G) both in vitro and in human cells. In addition, Cas12h1 utilizes a distinct activation mechanism that the lid motif undergoes a "flexible to stable" transition to expose the catalytic site to the substrate. A high-fidelity nucleic acid detector, Cas12h1 hf , was developed through rational engineering, which distinguishes single-base mismatches and retains comparable on-target activities. Our results shed light on the molecular mechanisms underlying Cas12h1 nickase, improve the understanding of type V Cas effectors, and expand the CRISPR toolbox for genome editing and molecular diagnosis.
Organizational Affiliation:
Key Laboratory of Microbial Pathogenesis and Interventions of Fujian Province University, the Key Laboratory of Innate Immune Biology of Fujian Province, Biomedical Research Center of South China, Key Laboratory of OptoElectronic Science and Technology for Medicine of the Ministry of Education, College of Life Sciences, Fujian Normal University, Fuzhou, 350117, China.