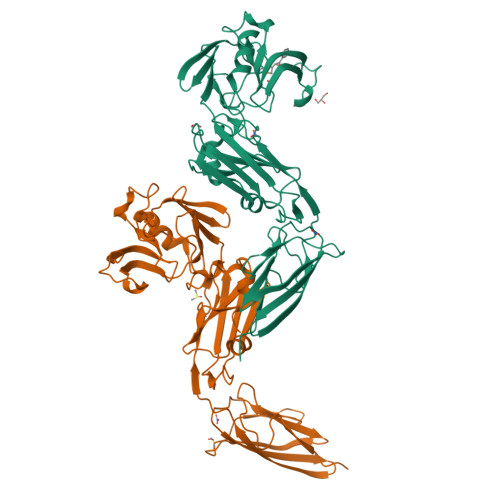
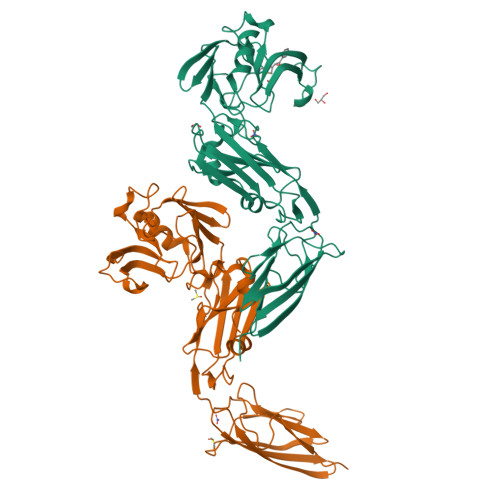
alpha beta , alpha ' beta '-Diepoxyketones are mechanism-based inhibitors of nucleophilic cysteine enzymes.
de Munnik, M., Lithgow, J., Brewitz, L., Christensen, K.E., Bates, R.H., Rodriguez-Miquel, B., Schofield, C.J.(2023) Chem Commun (Camb) 59: 12859-12862
- PubMed: 37815791
- DOI: https://doi.org/10.1039/d3cc02932h
- Primary Citation of Related Structures:
8BK3 - PubMed Abstract:
Epoxides are an established class of electrophilic alkylating agents that react with nucleophilic protein residues. We report αβ,α'β'-diepoxyketones (DEKs) as a new type of mechanism-based inhibitors of nucleophilic cysteine enzymes. Studies with the L,D-transpeptidase Ldt Mt2 from Mycobacterium tuberculosis and the main protease from SARS-CoV-2 (M pro ) reveal that following epoxide ring opening by a nucleophilic cysteine, further reactions can occur, leading to irreversible alkylation.
Organizational Affiliation:
Chemistry Research Laboratory, Department of Chemistry and the Ineos Oxford Institute of Antimicrobial Research, University of Oxford, 12 Mansfield Road, Oxford, OX1 3TA, UK. christopher.schofield@chem.ox.ac.uk.