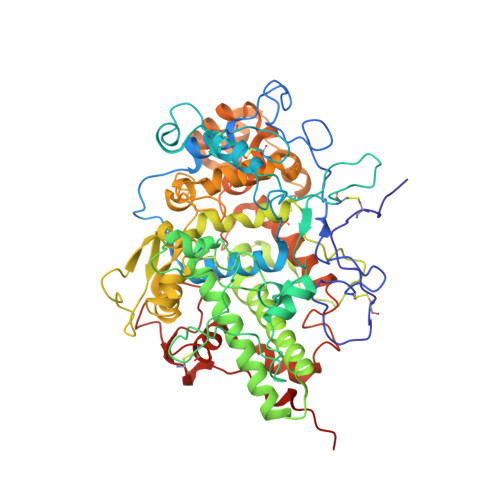
Potassium-induced partial inhibition of lactoperoxidase: structure of the complex of lactoperoxidase with potassium ion at 2.20 angstrom resolution.
Singh, P.K., Pandey, S., Rani, C., Ahmad, N., Viswanathan, V., Sharma, P., Kaur, P., Sharma, S., Singh, T.P.(2021) J Biol Inorg Chem 26: 149-159
- PubMed: 33427997
- DOI: https://doi.org/10.1007/s00775-020-01844-6
- Primary Citation of Related Structures:
6L32, 6L5G, 7C73, 7C74, 7C75, 7D52, 7DMR - PubMed Abstract:
Lactoperoxidase, a heme-containing glycoprotein, catalyzes the oxidation of thiocyanate by hydrogen peroxide into hypothiocyanite which acts as an antibacterial agent. The prosthetic heme moiety is attached to the protein through two ester linkages via Glu258 and Asp108. In lactoperoxidase, the substrate-binding site is formed on the distal heme side. To study the effect of physiologically important potassium ion on the structure and function of lactoperoxidase, the fresh protein samples were isolated from yak (Bos grunniens) colostrum and purified to homogeneity. The biochemical studies with potassium fluoride showed a significant reduction in the catalytic activity. Lactoperoxidase was crystallized using 200 mM ammonium nitrate and 20% PEG-3350 at pH 6.0. The crystals of LPO were soaked in the solution of potassium fluoride and used for the X-ray intensity data collection. Structure determination at 2.20 Å resolution revealed the presence of a potassium ion in the distal heme cavity. Structure determination further revealed that the propionic chain attached to pyrrole ring C of the heme moiety, was disordered into two components each having an occupancy of 0.5. One component occupied a position similar to the normally observed position of propionic chain while the second component was found in the distal heme cavity. The potassium ion in the distal heme cavity formed five coordinate bonds with two oxygen atoms of propionic moiety, N ε2 atom of His109 and two oxygen atoms of water molecules. The presence of potassium ion in the distal heme cavity hampered the catalytic activity of lactoperoxidase.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, Ansari Nagar, New Delhi, 110 029, India.