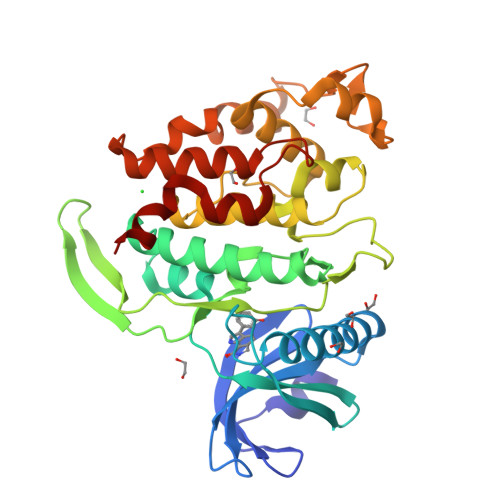
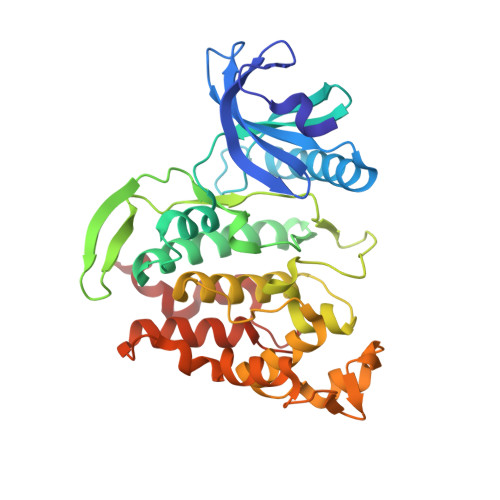
Molecular structures of cdc2-like kinases in complex with a new inhibitor chemotype.
Walter, A., Chaikuad, A., Helmer, R., Loaec, N., Preu, L., Ott, I., Knapp, S., Meijer, L., Kunick, C.(2018) PLoS One 13: e0196761-e0196761
- PubMed: 29723265
- DOI: https://doi.org/10.1371/journal.pone.0196761
- Primary Citation of Related Structures:
6FT7, 6FT8, 6FT9 - PubMed Abstract:
Cdc2-like kinases (CLKs) represent a family of serine-threonine kinases involved in the regulation of splicing by phosphorylation of SR-proteins and other splicing factors. Although compounds acting against CLKs have been described, only a few show selectivity against dual-specificity tyrosine phosphorylation regulated-kinases (DYRKs). We here report a novel CLK inhibitor family based on a 6,7-dihydropyrrolo[3,4-g]indol-8(1H)-one core scaffold. Within the series, 3-(3-chlorophenyl)-6,7-dihydropyrrolo[3,4-g]indol-8(1H)-one (KuWal151) was identified as inhibitor of CLK1, CLK2 and CLK4 with a high selectivity margin towards DYRK kinases. The compound displayed a potent antiproliferative activity in an array of cultured cancer cell lines. The X-ray structure analyses of three members of the new compound class co-crystallized with CLK proteins corroborated a molecular binding mode predicted by docking studies.
Organizational Affiliation:
Institut für Medizinische und Pharmazeutische Chemie, Technische Universität Braunschweig, Braunschweig, Germany.