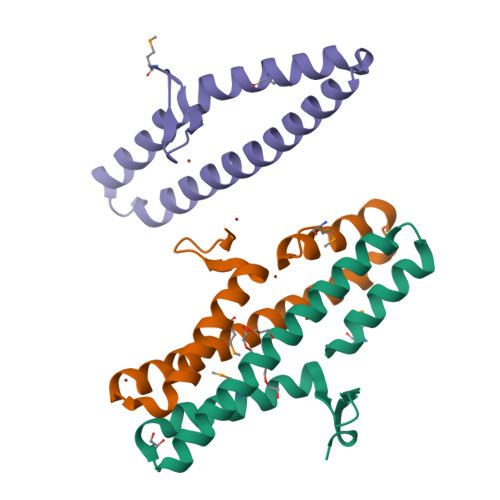
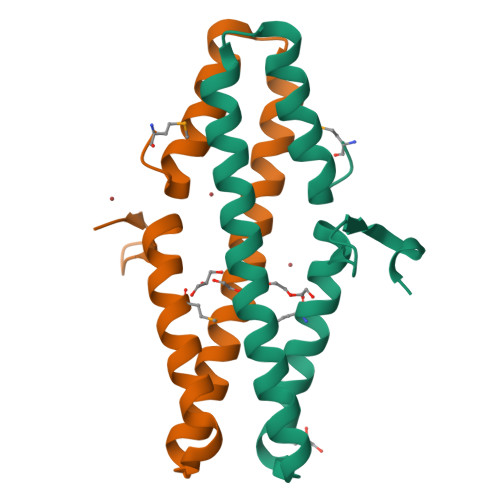
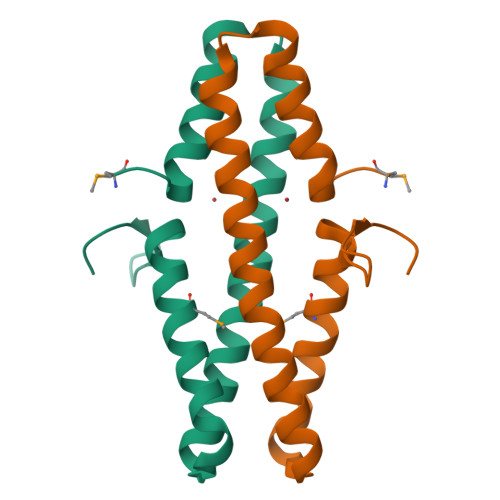
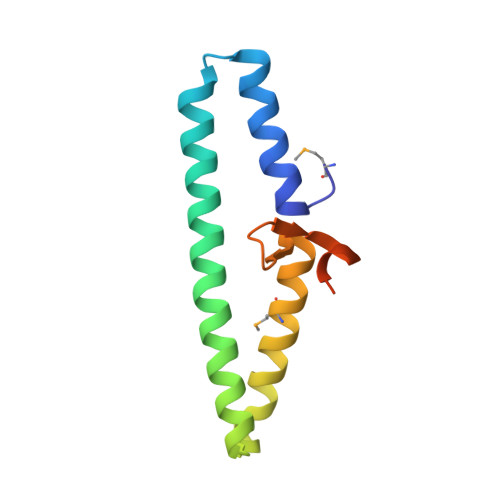
Disabling a Type I-E CRISPR-Cas Nuclease with a Bacteriophage-Encoded Anti-CRISPR Protein.
Pawluk, A., Shah, M., Mejdani, M., Calmettes, C., Moraes, T.F., Davidson, A.R., Maxwell, K.L.(2017) mBio 8
- PubMed: 29233895
- DOI: https://doi.org/10.1128/mBio.01751-17
- Primary Citation of Related Structures:
6ARZ, 6AS3, 6AS4 - PubMed Abstract:
CRISPR (clustered regularly interspaced short palindromic repeat)-Cas adaptive immune systems are prevalent defense mechanisms in bacteria and archaea. They provide sequence-specific detection and neutralization of foreign nucleic acids such as bacteriophages and plasmids. One mechanism by which phages and other mobile genetic elements are able to overcome the CRISPR-Cas system is through the expression of anti-CRISPR proteins. Over 20 different families of anti-CRISPR proteins have been described, each of which inhibits a particular type of CRISPR-Cas system. In this work, we determined the structure of type I-E anti-CRISPR protein AcrE1 by X-ray crystallography. We show that AcrE1 binds to the CRISPR-associated helicase/nuclease Cas3 and that the C-terminal region of the anti-CRISPR protein is important for its inhibitory activity. We further show that AcrE1 can convert the endogenous type I-E CRISPR system into a programmable transcriptional repressor. IMPORTANCE The CRISPR-Cas immune system provides bacteria with resistance to invasion by potentially harmful viruses, plasmids, and other foreign mobile genetic elements. This study presents the first structural and mechanistic insight into a phage-encoded protein that inactivates the type I-E CRISPR-Cas system in Pseudomonas aeruginosa The interaction of this anti-CRISPR protein with the CRISPR-associated helicase/nuclease proteins Cas3 shuts down the CRISPR-Cas system and protects phages carrying this gene from destruction. This interaction also allows the repurposing of the endogenous type I-E CRISPR system into a programmable transcriptional repressor, providing a new biotechnological tool for genetic studies of bacteria encoding this type I-E CRISPR-Cas system.
Organizational Affiliation:
Department of Biochemistry, University of Toronto, Toronto, Ontario, Canada.