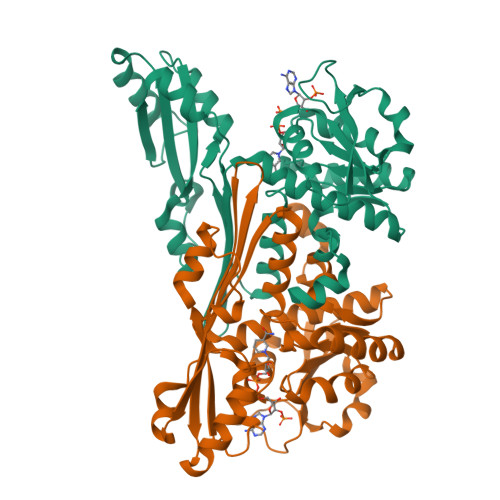
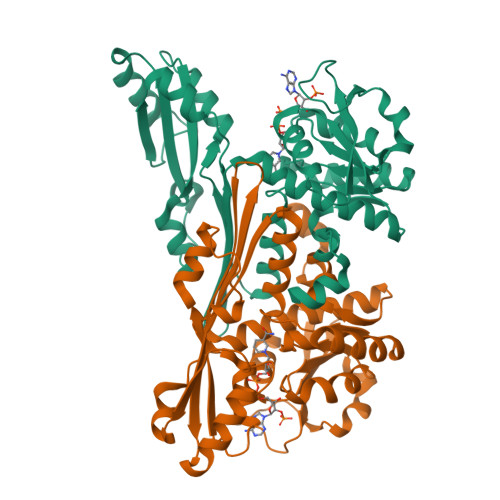
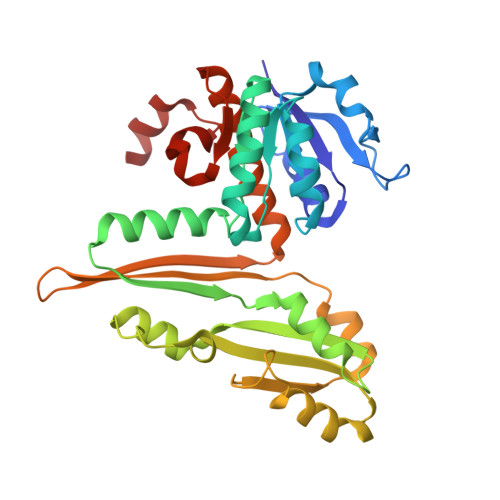
Single-biocatalyst synthesis of enantiopure D-arylalanines exploiting an engineered D-amino acid dehydrogenase
Parmeggiani, F., Ahmed, S.T., Thompson, M.P., Weise, N.J., Galman, J.L., Gahloth, D., Dunstan, M.S., Leys, D., Turner, N.J.To be published.