Determination of Kinetics and the Crystal Structure of a Novel Type 2 Isopentenyl Diphosphate: Dimethylallyl Diphosphate Isomerase from Streptococcus pneumoniae.
de Ruyck, J., Janczak, M.W., Neti, S.S., Rothman, S.C., Schubert, H.L., Cornish, R.M., Matagne, A., Wouters, J., Poulter, C.D.(2014) Chembiochem 15: 1452-1458
- PubMed: 24910111
- DOI: https://doi.org/10.1002/cbic.201402046
- Primary Citation of Related Structures:
4N02 - PubMed Abstract:
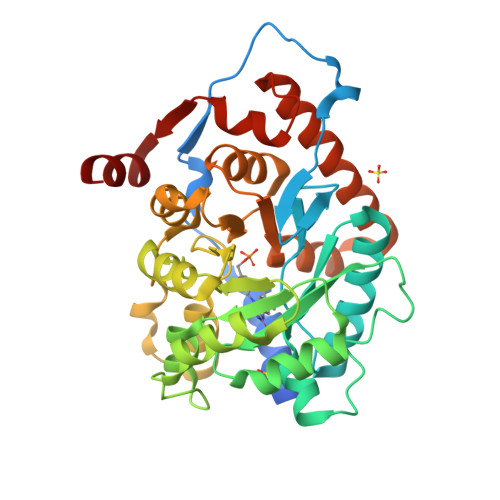
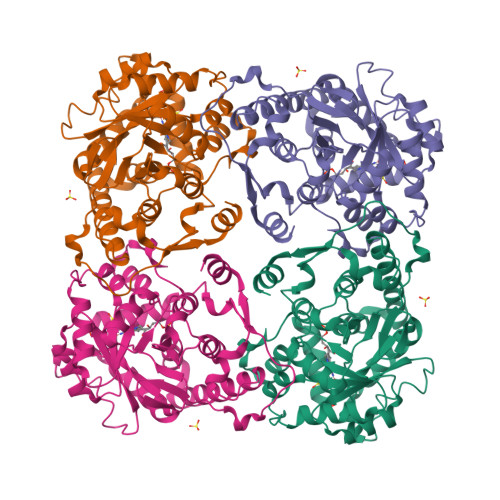
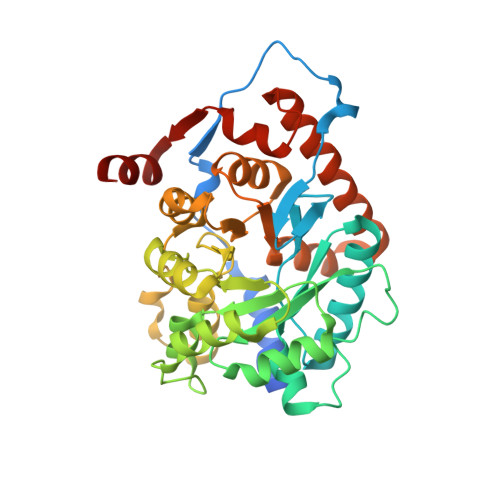
Isopentenyl diphosphate isomerase (IDI) is a key enzyme in the isoprenoid biosynthetic pathway and is required for all organisms that synthesize isoprenoid metabolites from mevalonate. Type 1 IDI (IDI-1) is a metalloprotein that is found in eukaryotes, whereas the type 2 isoform (IDI-2) is a flavoenzyme found in bacteria that is completely absent from human. IDI-2 from the pathogenic bacterium Streptococcus pneumoniae was recombinantly expressed in Escherichia coli. Steady-state kinetic studies of the enzyme indicated that FMNH2 (KM =0.3 μM) bound before isopentenyl diphosphate (KM =40 μM) in an ordered binding mechanism. An X-ray crystal structure at 1.4 Å resolution was obtained for the holoenzyme in the closed conformation with a reduced flavin cofactor and two sulfate ions in the active site. These results helped to further approach the enzymatic mechanism of IDI-2 and, thus, open new possibilities for the rational design of antibacterial compounds against sequence-similar and structure-related pathogens such as Enterococcus faecalis or Staphylococcus aureus.
Organizational Affiliation:
Department of Chemistry, University of Utah, 315 South 1400 East RM 2020, Salt Lake City, Utah 84112 (USA); Department of Chemistry, UNamur, 61 rue de Bruxelles, 5000 Namur (Belgium).