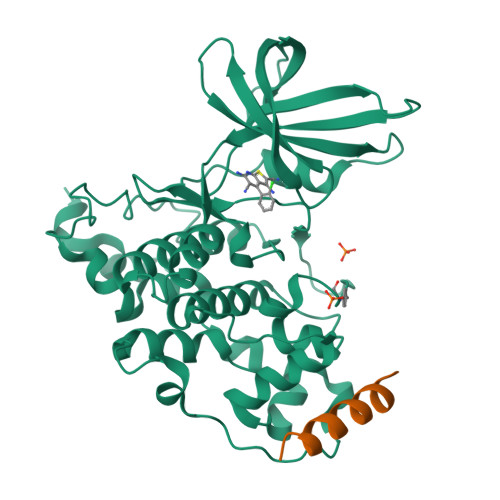
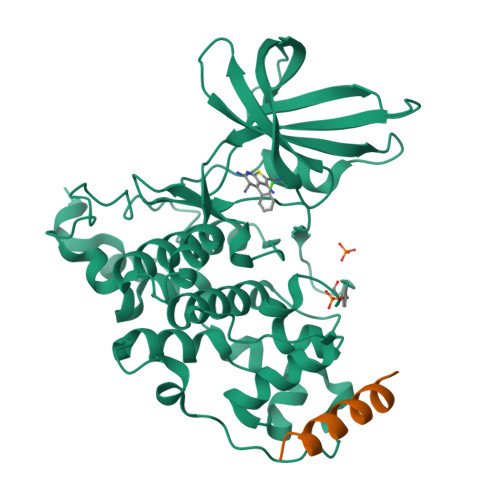
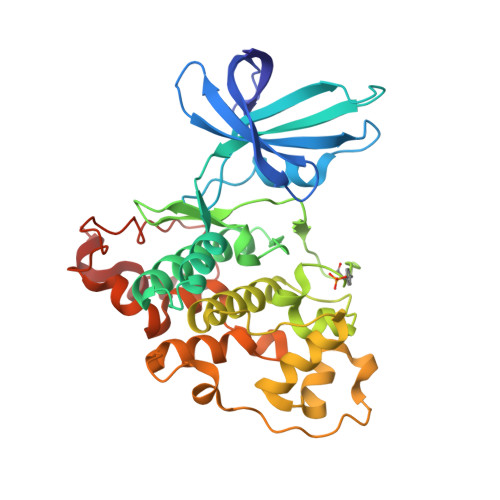
3,6-Diamino-4-(2-Halophenyl)-2-Benzoylthieno(2,3-B) Pyridine-5-Carbonitriles are Selective Inhibitors of Plasmodium Falciparum Glycogen Synthase Kinase-3 (Pfgsk-3)
Fugel, W., Oberholzer, A.E., Gschloessl, B., Dzikowski, R., Pressburger, N., Preu, L., Pearl, L.H., Baratte, B., Ratin, M., Okun, I., Doerig, C., Kruggel, S., Lemcke, T., Meijer, L., Kunick, C.(2013) J Med Chem 56: 264
- PubMed: 23214499
- DOI: https://doi.org/10.1021/jm301575n
- Primary Citation of Related Structures:
3ZDI - PubMed Abstract:
Plasmodium falciparum is the infective agent responsible for malaria tropica. The glycogen synthase kinase-3 of the parasite (PfGSK-3) was suggested as a potential biological target for novel antimalarial drugs. Starting from hit structures identified in a high-throughput screening campaign, 3,6-diamino-4-(2-halophenyl)-2-benzoylthieno[2,3-b]pyridine-5-carbonitriles were discovered as a new class of PfGSK-3 inhibitors. Being less active on GSK-3 homologues of other species, the title compounds showed selectivity in favor of PfGSK-3. Taking into account the X-ray structure of a related molecule in complex with human GSK-3 (HsGSK-3), a model was computed for the comparison of inhibitor complexes with the plasmodial and human enzymes. It was found that subtle differences in the ATP-binding pockets are responsible for the observed PfGSK-3 vs HsGSK-3 selectivity. Representatives of the title compound class exhibited micromolar IC₅₀ values against P. falciparum erythrocyte stage parasites. These results suggest that inhibitors of PfGSK-3 could be developed as potential antimalarial drugs.
Organizational Affiliation:
Institut für Medizinische und Pharmazeutische Chemie, Technische Universität Braunschweig, Beethovenstrasse 55, 38106 Braunschweig, Germany.