Fused bicyclic heteroarylpiperazine-substituted l-prolylthiazolidines as highly potent DPP-4 inhibitors lacking the electrophilic nitrile group
Yoshida, T., Akahoshi, F., Sakashita, H., Sonda, S., Takeuchi, M., Tanaka, Y., Nabeno, M., Kishida, H., Miyaguchi, I., Hayashi, Y.(2012) Bioorg Med Chem 20: 5033-5041
- PubMed: 22824762
- DOI: https://doi.org/10.1016/j.bmc.2012.06.033
- Primary Citation of Related Structures:
3VJM - PubMed Abstract:
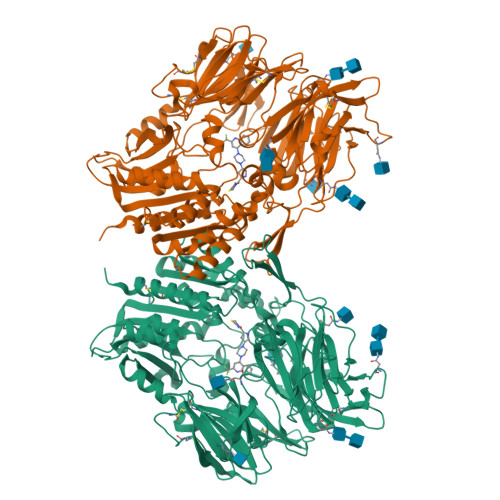
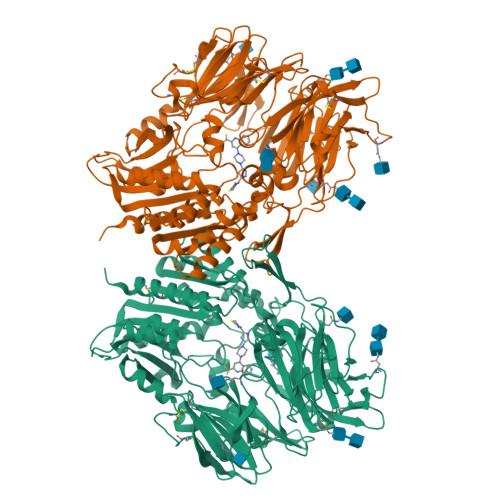
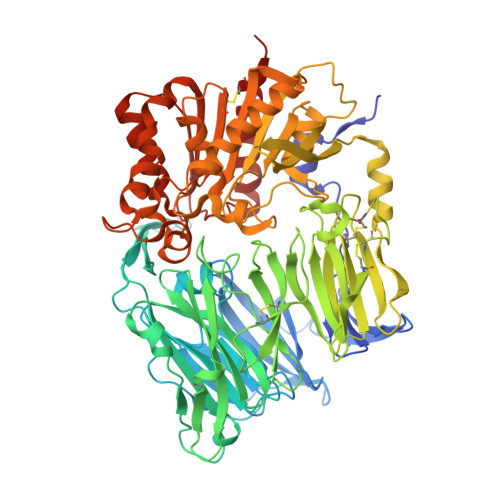
Hypoglycemic agents with a mechanism of depeptidyl peptidase IV (DPP-4) inhibition are suitable for once daily oral dosing. It is difficult to strike a balance between inhibitory activity and duration of action in plasma for inhibitors bearing an electrophilic nitrile group. We explored fused bicyclic heteroarylpiperazine substituted at the γ-position of the proline structure in the investigation of L-prolylthiazolidines lacking the electrophilic nitrile. Among them, 2-trifluoroquinolyl compound 8g is the most potent, long-lasting DPP-4 inhibitor (IC(50) = 0.37 nmol/L) with high selectivity against other related peptidases. X-ray crystal structure determination of 8g indicates that CH-π interactions generated between the quinolyl ring and the guanidinyl group of Arg358 enhances the DPP-4 inhibitory activity and selectivity.
Organizational Affiliation:
Research Division, Mitsubishi Tanabe Pharma Corporation, 2-2-50, Kawagishi, Toda-shi, Saitama 335-8505, Japan.