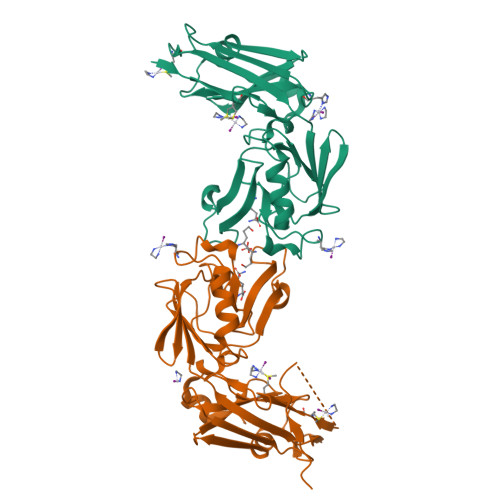
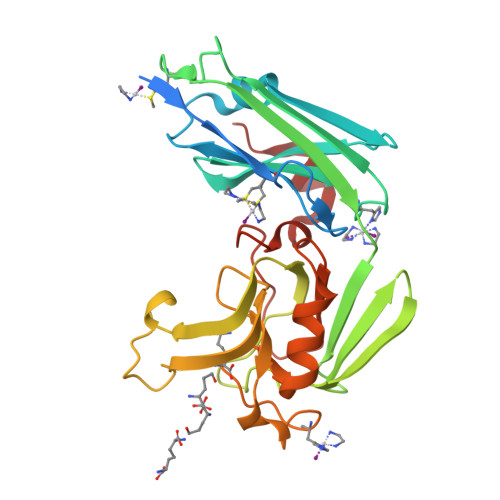
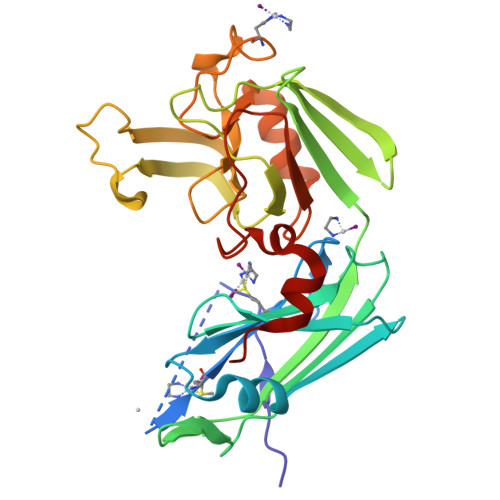
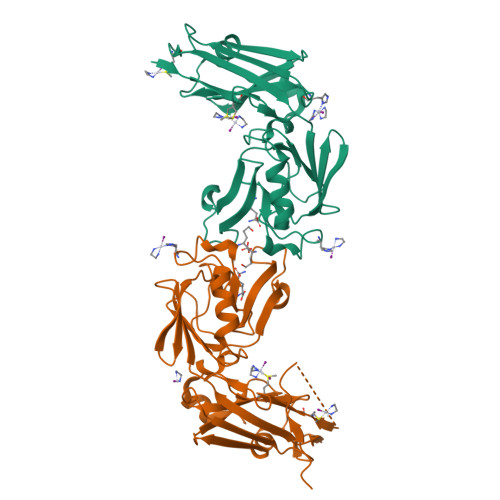
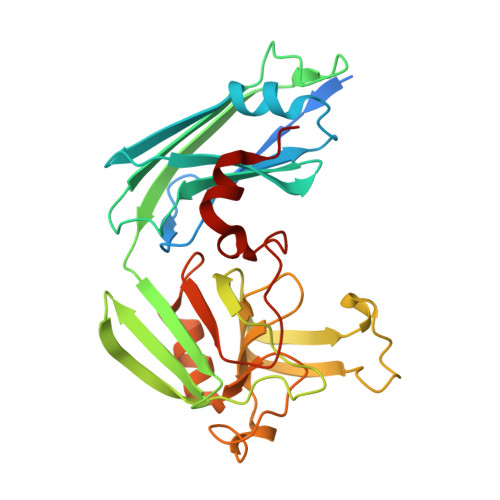
Targeting the Cell Wall of Mycobacterium tuberculosis: Structure and Mechanism of L,D-Transpeptidase 2.
Erdemli, S.B., Gupta, R., Bishai, W.R., Lamichhane, G., Amzel, L.M., Bianchet, M.A.(2012) Structure 20: 2103-2115
- PubMed: 23103390
- DOI: https://doi.org/10.1016/j.str.2012.09.016
- Primary Citation of Related Structures:
3TUR, 3TX4, 3U1P, 3VAE - PubMed Abstract:
With multidrug-resistant cases of tuberculosis increasing globally, better antibiotic drugs and novel drug targets are becoming an urgent need. Traditional β-lactam antibiotics that inhibit D,D-transpeptidases are not effective against mycobacteria, in part because mycobacteria rely mostly on L,D-transpeptidases for biosynthesis and maintenance of their peptidoglycan layer. This reliance plays a major role in drug resistance and persistence of Mycobacterium tuberculosis (Mtb) infections. The crystal structure at 1.7 Å resolution of the Mtb L,D-transpeptidase Ldt(Mt2) containing a bound peptidoglycan fragment, reported here, provides information about catalytic site organization as well as substrate recognition by the enzyme. Based on our structural, kinetic, and calorimetric data, we propose a catalytic mechanism for Ldt(Mt2) in which both acyl-acceptor and acyl-donor substrates reach the catalytic site from the same, rather than different, entrances. Together, this information provides vital insights to facilitate development of drugs targeting this validated yet unexploited enzyme.
Organizational Affiliation:
Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.