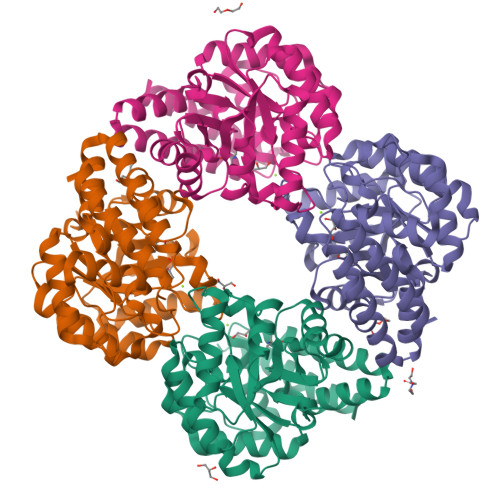
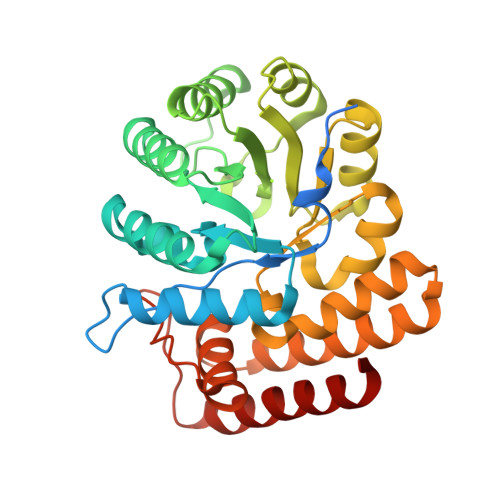
Elucidation of a sialic acid metabolism pathway in mucus-foraging Ruminococcus gnavus unravels mechanisms of bacterial adaptation to the gut.
Bell, A., Brunt, J., Crost, E., Vaux, L., Nepravishta, R., Owen, C.D., Latousakis, D., Xiao, A., Li, W., Chen, X., Walsh, M.A., Claesen, J., Angulo, J., Thomas, G.H., Juge, N.(2019) Nat Microbiol 4: 2393-2404
- PubMed: 31636419
- DOI: https://doi.org/10.1038/s41564-019-0590-7
- Primary Citation of Related Structures:
6RAB, 6RB7, 6RD1 - PubMed Abstract:
Sialic acid (N-acetylneuraminic acid (Neu5Ac)) is commonly found in the terminal location of colonic mucin glycans where it is a much-coveted nutrient for gut bacteria, including Ruminococcus gnavus. R. gnavus is part of the healthy gut microbiota in humans, but it is disproportionately represented in diseases. There is therefore a need to understand the molecular mechanisms that underpin the adaptation of R. gnavus to the gut. Previous in vitro research has demonstrated that the mucin-glycan-foraging strategy of R. gnavus is strain dependent and is associated with the expression of an intramolecular trans-sialidase, which releases 2,7-anhydro-Neu5Ac, rather than Neu5Ac, from mucins. Here, we unravelled the metabolism pathway of 2,7-anhydro-Neu5Ac in R. gnavus that is underpinned by the exquisite specificity of the sialic transporter for 2,7-anhydro-Neu5Ac and by the action of an oxidoreductase that converts 2,7-anhydro-Neu5Ac into Neu5Ac, which then becomes a substrate of a Neu5Ac-specific aldolase. Having generated an R. gnavus nan-cluster deletion mutant that lost the ability to grow on sialylated substrates, we showed that-in gnotobiotic mice colonized with R. gnavus wild-type (WT) and mutant strains-the fitness of the nan mutant was significantly impaired, with a reduced ability to colonize the mucus layer. Overall, we revealed a unique sialic acid pathway in bacteria that has important implications for the spatial adaptation of mucin-foraging gut symbionts in health and disease.
Organizational Affiliation:
The Gut Microbes and Health Institute Strategic Programme, Quadram Institute Bioscience, Norwich Research Park, Norwich, UK.