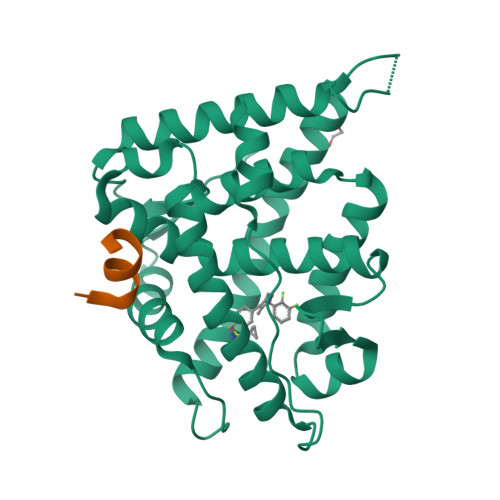
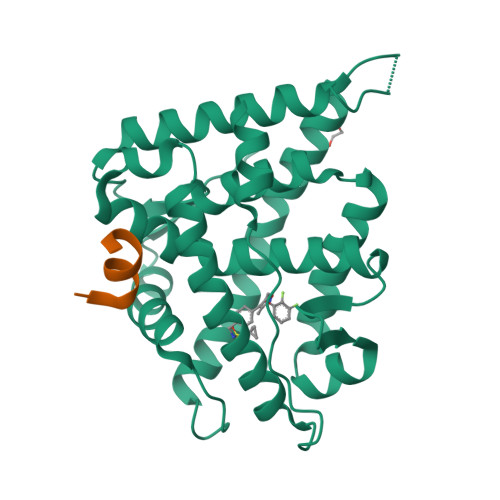
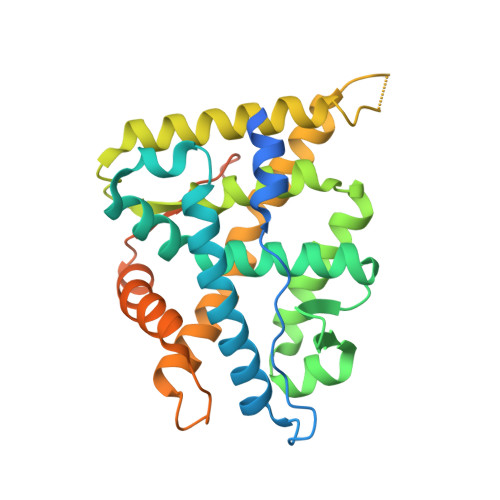
Structure-Based Drug Design of Mineralocorticoid Receptor Antagonists to Explore Oxosteroid Receptor Selectivity.
Nordqvist, A., O'Mahony, G., Friden-Saxin, M., Fredenwall, M., Hogner, A., Granberg, K.L., Aagaard, A., Backstrom, S., Gunnarsson, A., Kaminski, T., Xue, Y., Dellsen, A., Hansson, E., Hansson, P., Ivarsson, I., Karlsson, U., Bamberg, K., Hermansson, M., Georgsson, J., Lindmark, B., Edman, K.(2017) ChemMedChem 12: 50-65
- PubMed: 27897427
- DOI: https://doi.org/10.1002/cmdc.201600529
- Primary Citation of Related Structures:
5L7E, 5L7G, 5L7H - PubMed Abstract:
The mineralocorticoid receptor (MR) is a nuclear hormone receptor involved in the regulation of body fluid and electrolyte homeostasis. In this study we explore selectivity triggers for a series of nonsteroidal MR antagonists to improve selectivity over other members of the oxosteroid receptor family. A biaryl sulfonamide compound was identified in a high-throughput screening (HTS) campaign. The compound bound to MR with pK i =6.6, but displayed poor selectivity over the glucocorticoid receptor (GR) and the progesterone receptor (PR). Following X-ray crystallography of MR in complex with the HTS hit, a compound library was designed that explored an induced-fit hypothesis that required movement of the Met852 side chain. An improvement in MR selectivity of 11- to 79-fold over PR and 23- to 234-fold over GR was obtained. Given the U-shaped binding conformation, macrocyclizations were explored, yielding a macrocycle that bound to MR with pK i =7.3. Two protein-ligand X-ray structures were determined, confirming the hypothesized binding mode for the designed compounds.
Organizational Affiliation:
Cardiovascular and Metabolic Diseases, Innovative Medicines and Early Development Biotech Unit, AstraZeneca, Pepparedsleden 1, Mölndal, 43183, Sweden.