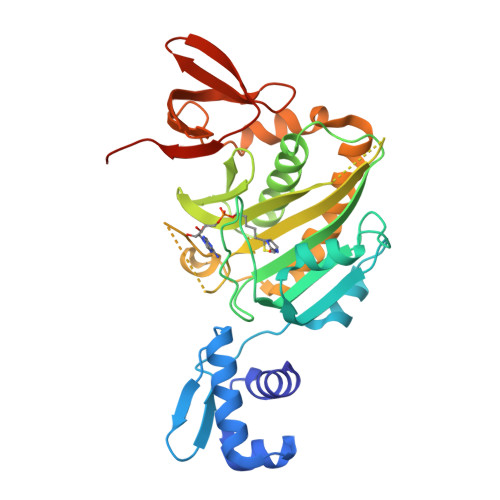
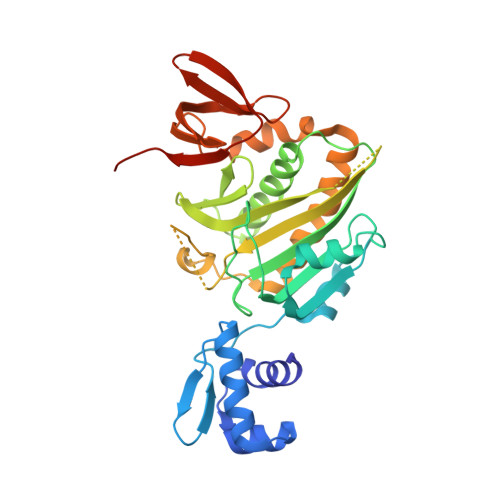
Allosteric Coupling via Distant Disorder-to-Order Transitions.
Eginton, C., Cressman, W.J., Bachas, S., Wade, H., Beckett, D.(2015) J Mol Biology 427: 1695-1704
- PubMed: 25746672
- DOI: https://doi.org/10.1016/j.jmb.2015.02.021
- Primary Citation of Related Structures:
4WF2 - PubMed Abstract:
Intrinsic disorder provides a means of maximizing allosteric coupling in proteins. However, the mechanisms by which the disorder functions in allostery remain to be elucidated. Small ligand, bio-5'-AMP, binding and dimerization of the Escherichia coli biotin repressor are allosterically coupled. Folding of a disordered loop in the allosteric effector binding site is required to realize the full coupling free energy of -4.0 ± 0.3 kcal/mol observed in the wild-type protein. Alanine substitution of a glycine residue on the dimerization surface that does not directly contribute to the dimerization interface completely abolishes this coupling. In this work, the structure of this variant, solved by X-ray crystallography, reveals a monomeric corepressor-bound protein. In the structure loops, neither of which contains the alanine substitution, on both the dimerization and effector binding surfaces that are folded in the corepressor-bound wild-type protein are disordered. The structural data combined with functional measurements indicate that allosteric coupling between ligand binding and dimerization in BirA (E. coli biotin repressor/biotin protein ligase) is achieved via reciprocal communication of disorder-to-order transitions on two distant functional surfaces.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Maryland, College Park, MD 20742, USA.