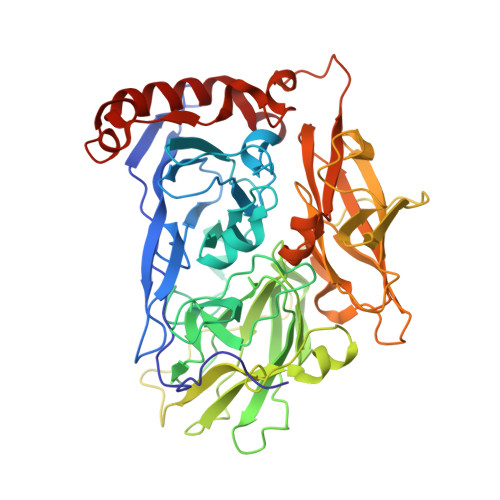
X-ray analysis of bilirubin oxidase from Myrothecium verrucaria at 2.3 A resolution using a twinned crystal
Mizutani, K., Toyoda, M., Sagara, K., Takahashi, N., Sato, A., Kamitaka, Y., Tsujimura, S., Nakanishi, Y., Sugiura, T., Yamaguchi, S., Kano, K., Mikami, B.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 765-770
- PubMed: 20606269
- DOI: https://doi.org/10.1107/S1744309110018828
- Primary Citation of Related Structures:
3ABG - PubMed Abstract:
Bilirubin oxidase (BOD), a multicopper oxidase found in Myrothecium verrucaria, catalyzes the oxidation of bilirubin to biliverdin. Oxygen is the electron acceptor and is reduced to water. BOD is used for diagnostic analysis of bilirubin in serum and has attracted considerable attention as an enzymatic catalyst for the cathode of biofuel cells that work under neutral conditions. Here, the crystal structure of BOD is reported for the first time. Blue bipyramid-shaped crystals of BOD obtained in 2-methyl-2,4-pentanediol (MPD) and ammonium sulfate solution were merohedrally twinned in space group P6(3). Structure determination was achieved by the single anomalous diffraction (SAD) method using the anomalous diffraction of Cu atoms and synchrotron radiation and twin refinement was performed in the resolution range 33-2.3 A. The overall organization of BOD is almost the same as that of other multicopper oxidases: the protein is folded into three domains and a total of four copper-binding sites are found in domains 1 and 3. Although the four copper-binding sites were almost identical to those of other multicopper oxidases, the hydrophilic Asn residue (at the same position as a hydrophobic residue such as Leu in other multicopper oxidases) very close to the type I copper might contribute to the characteristically high redox potential of BOD.
Organizational Affiliation:
Laboratory of Applied Structural Biology, Division of Applied Life Sciences, The Graduate School of Agriculture, Kyoto University, Uji, Kyoto 611-0011, Japan. [email protected]