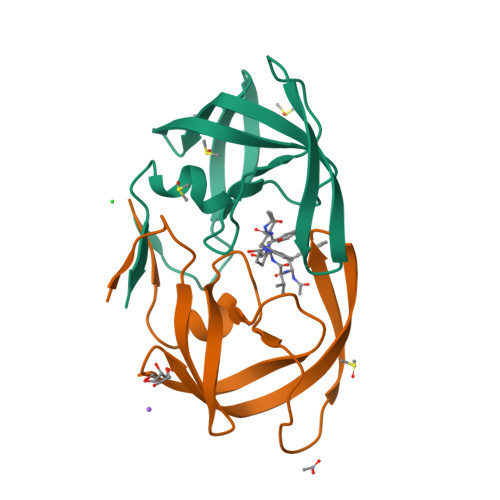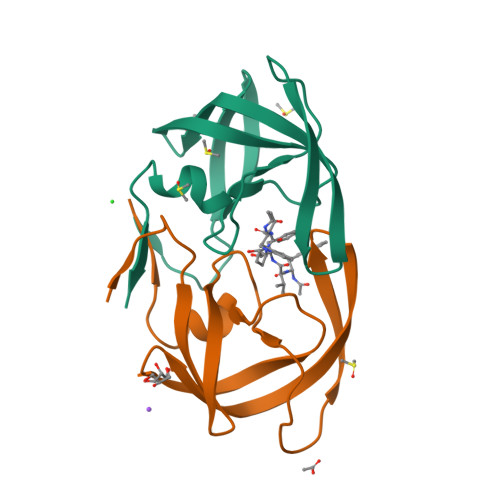A potent HIV protease inhibitor identified in an epimeric mixture by high-resolution protein crystallography.
Geremia, S., Demitri, N., Wuerges, J., Benedetti, F., Berti, F., Tell, G., Randaccio, L.(2006) ChemMedChem 1: 186-188
- PubMed: 16892350
- DOI: https://doi.org/10.1002/cmdc.200500063
- Primary Citation of Related Structures:
2A1E
Organizational Affiliation:
Department of Chemical Sciences and Centre of Excellence in Biocrystallography, University of Trieste, Via L. Giorgieri 1, 34127 Trieste, Italy. geremia@univ.trieste.it