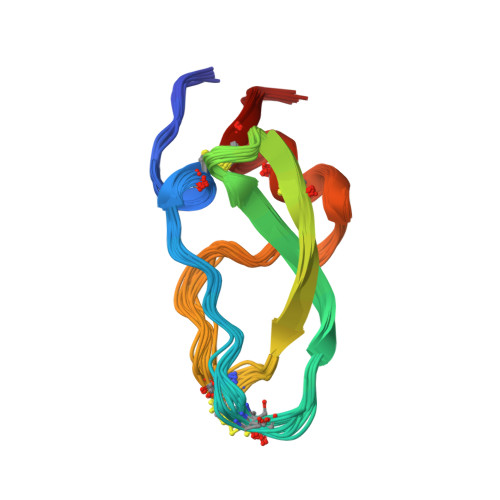
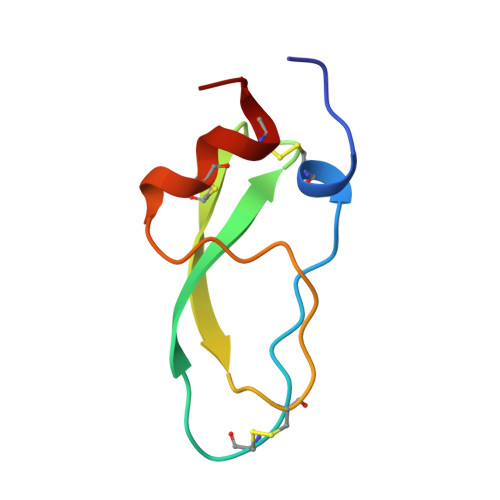
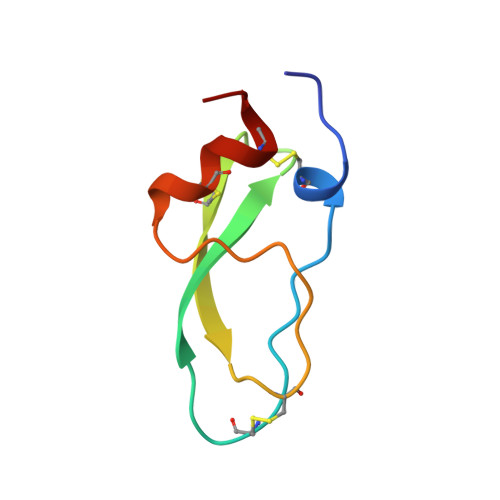
Conformational and functional variability supported by the BPTI fold: solution structure of the Ca2+ channel blocker calcicludine.
Gilquin, B., Lecoq, A., Desne, F., Guenneugues, M., Zinn-Justin, S., Menez, A.(1999) Proteins 34: 520-532
- PubMed: 10081964
- Primary Citation of Related Structures:
1BF0 - PubMed Abstract:
Calcicludine, a 60-amino acid protein isolated from the green mamba venom, has been recently identified as blocking a large set (i.e., L-, N- and P-type) of Ca2+ channels. The three-dimensional structure of calcicludine has been determined by NMR and molecular modeling using a data set of 723 unambiguous and 265 ambiguous distance restraints, as 33 phi and 13 chi1 dihedral angle restraints. Analysis of the 15 final structures (backbone root-mean-square deviation = 0.6 A) shows that calcicludine adopts the Kunitz-type protease inhibitor fold. Its three-dimensional structure is similar to that of snake K+ channel blockers dendrotoxins. Conformational differences with protease inhibitors and dendrotoxins are localized in the 3(10) helix and loop 1 (segments 1-7 and 10-19), the extremity of the beta-hairpin (segment 27-30), and loop 2 (segment 39-44). These regions correspond to the functional sites of bovine pancreatic trypsin inhibitor (BPTI) and dendrotoxins. The positioning of the N-terminal segment 1-7 relative to the rest of the protein is characteristic of calcicludine. The involvement of this segment and the positively charged K31 at the tip of the beta-hairpin in the biological activity of calcicludine is discussed.
Organizational Affiliation:
CEA, Département d'Ingénierie et d'Etudes des Protéines, Gif-sur-Yvette, France.